_25.png)
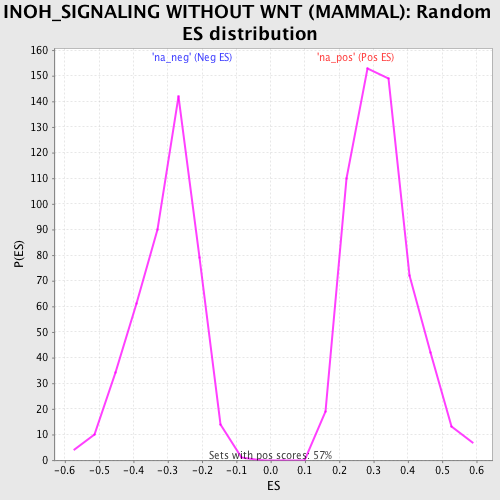
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | INOH_SIGNALING WITHOUT WNT (MAMMAL) |
| Enrichment Score (ES) | 0.6535035 |
| Normalized Enrichment Score (NES) | 2.0408657 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.004736345 |
| FWER p-Value | 0.062 |
_25.png)
| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DVL2 | 380 | 8.902 | 0.0723 | Yes | ||
| 2 | LEF1 | 400 | 8.761 | 0.1597 | Yes | ||
| 3 | BCL9 | 488 | 8.237 | 0.2386 | Yes | ||
| 4 | TCF7L2 | 546 | 7.944 | 0.3160 | Yes | ||
| 5 | TCF7 | 1011 | 6.319 | 0.3585 | Yes | ||
| 6 | AXIN1 | 1475 | 5.258 | 0.3903 | Yes | ||
| 7 | TLE4 | 1595 | 5.061 | 0.4358 | Yes | ||
| 8 | TLE1 | 1674 | 4.931 | 0.4819 | Yes | ||
| 9 | TLE3 | 1727 | 4.860 | 0.5285 | Yes | ||
| 10 | CREBBP | 2209 | 4.217 | 0.5490 | Yes | ||
| 11 | NLK | 2430 | 3.973 | 0.5790 | Yes | ||
| 12 | DVL1 | 2689 | 3.713 | 0.6046 | Yes | ||
| 13 | GSK3B | 2980 | 3.451 | 0.6261 | Yes | ||
| 14 | FRAT2 | 3115 | 3.330 | 0.6535 | Yes | ||
| 15 | APC | 4019 | 2.635 | 0.6388 | No | ||
| 16 | SMAD4 | 4759 | 2.167 | 0.6270 | No | ||
| 17 | CUL1 | 4951 | 2.070 | 0.6391 | No | ||
| 18 | TLE2 | 5275 | 1.886 | 0.6433 | No | ||
| 19 | TLE6 | 6664 | 1.226 | 0.5924 | No | ||
| 20 | AXIN2 | 6861 | 1.146 | 0.5950 | No | ||
| 21 | DVL3 | 8296 | 0.541 | 0.5350 | No | ||
| 22 | CTNNBIP1 | 10901 | -0.548 | 0.4218 | No | ||
| 23 | CTNNB1 | 11600 | -0.784 | 0.3978 | No | ||
| 24 | TCF3 | 17265 | -2.559 | 0.1652 | No | ||
| 25 | PYGO1 | 20564 | -4.798 | 0.0631 | No |